Inhalt
Prof. Maria Leptin
Institute for Genetics
+49 221 470 3401
+49 221 470 5264
The role of NLR proteins in pathogen recognition and defense
General background: innate immunity
Organisms need to be able to defend themselves against pathogens to survive. The ability of higher vertebrates to develop immunity has long been recognized, and the mechanisms are well understood. However, an ‘adaptive’ immune response takes several days to develop, and many pathogens can kill an organism within a much shorter period. The vital initial defense against pathogens therefore requires a faster response, which is mediated by a range of ‘innate’ immune mechanisms, many of which are shared by all animals, and some even by plants.
An important component of the innate immune response is the initial detection of the presence of pathogens and the triggering of the appropriate cellular reaction. The family of NLR proteins have been shown to be involved in the reaction to signals emanating from bacteria in the cytoplasm. In mammals a number of such molecules are known, but it is not clear how they deal with a large variety of different and evolving pathogens.
The work of our lab: pathogen defense mechanisms in the zebrafish
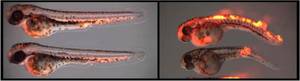
We use the zebrafish embryo to study innate immune mechanisms. The advantage of the zebrafish embryo is that it can be easily manipulated, physiological processes can be observed by live imaging, and above all, that its adaptive immune system is not yet active. We have discovered that the genome of the zebrafish contains more than 200 genes encoding fish-specific NLR proteins with both conserved and divergent domains. In addition, we have found a new NLR protein that is conserved among all vertebrates. We have shown that the divergent NLR proteins contain a previously unknown domain in their N-terminal part, the Fisna domain.
Project
The aim of the project will be to investigate how the novel NLR proteins are involved in pathogen recognition and contribute to innate immunity. We expect to find novel mechanisms responsible for the detection and elimination of bacteria. We will construct assay systems in which immune responses can be monitored in vivo by fluorescent detection systems. Our newly generated antibodies will be used to identify the interaction partners with which the NLR proteins associate to fulfil their functions. Protein complexes will be immunoprecipitated and analysed by mass spectrometry. Potential interaction partners will be studied by knocking down their function in the embryo using morpholinos. The work will be complemented by studying the biochemical properties of the proteins in cell culture and in vitro.
Generation of complex cell shapes and the role of localized mRNAs
Background: Tracheal cells
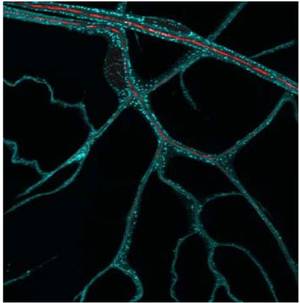
Many cells have highly polarized morphologies that serve specialized cell functions. We use the terminal cells of the Drosophila respiratory (tracheal) system to study the processes that generate complex cell shapes. Terminal cells, like neurons, are extensively branched. The mechanisms involved in establishing their cellular architecture, especially the oxygencarrying lumen, are poorly understood, as is also the case for related events in vertebrate tissues, in particular during the process of neo-angiogenesis as it occurs for example in tumor growth. Tracheal cell morphology depends on environ- mental signals that trigger events at sites distant from the cell body. We therefore assume that it depends on protein synthesis from locally stored RNA. Subcellular localization of mRNA and the local control of translation are indeed increasingly recognized as significant for polarized cellular functions However, general rules for the role of mRNA localization and the cis-regulatory elements that guide mRNAs to their destinations are yet to be discovered. Our work on tracheal cells has two aims: the understanding of complex cell morphology and the search for signals involved in mRNA localisation.
Approaches
We have developed an in vivo screening method for the identification of genes with asymmetrically localized RNA. A pilot screen for mRNAs localized in the branches of tracheal cells has shown that we can identify genes required for cell morphogenesis that have not been found by other methods. We will study these genes with a view to discovering the mechanisms by which branches grow in response to local cues and how an intracellular lumen is formed. For a full understanding of the processes we plan to discover further genes by conducting a full-scale screen. Results from testing genes from the pilot screen in other cell types have indicated that there must be cell type specific mRNA recognition and processing systems. To build a sequence database of mRNAs with tissue-specific polar distributions, the data obtained through screening tracheal cells in our own group will be enriched by input from collaborating laboratories who will use the insertion stocks we generate to screen other cell types. The combined datasets will be used for the bioinformatic identification of localisation signals.
Project
Students joining the lab have many choices for their research projects: - they may participate in the screen in order to discover new genes, among which they can then choose a subset for further studies - they may follow up initial results from the analysis of mutants and genes already under investigation in the lab. This will involve, among other aspects, the study of molecules involved in membrane trafficking and fusion. - they may use a combination of experimental and bioinformatic approaches to determine mRNA localisation signals. Methods will include live imaging of branch outgrowth and lumen formation, generation of mutants and sophisticated genetic engineering of flies, biochemistry and cell culture.